Evaluate Stability Across Resolution, Number of Neighbors, and Graph Type
Source:R/stability-based-parameter-assessment.R
get_resolution_importance.RdPerform a grid search over the resolution, number of neighbors and graph type.
get_resolution_importance(
embedding,
resolution,
n_neigh,
n_repetitions = 100,
seed_sequence = NULL,
clustering_method = 4,
graph_type = 0,
object_name = NULL,
ecs_thresh = 1,
ncores = 1
)Arguments
- embedding
The base embedding for the graph construction.
- resolution
A sequence of resolution values.
- n_neigh
A value or a sequence of number of neighbors used for graph construction.
- n_repetitions
The number of repetitions of applying the pipeline with different seeds; ignored if seed_sequence is provided by the user.
- seed_sequence
A custom seed sequence; if the value is NULL, the sequence will be built starting from 1 with a step of 100.
- clustering_method
An index or a list of indexes indicating which community detection algorithm will be used: Louvain (1), Louvain refined (2), SLM (3) or Leiden (4). More details can be found in the Seurat's
FindClustersfunction.- graph_type
Argument indicating whether the graph should be unweighted (0), weighted (1) or both (2).
- object_name
User specified string that uniquely describes the embedding characteristics.
- ecs_thresh
The ECS threshold used for merging similar clusterings.
- ncores
The number of parallel R instances that will run the code. If the value is set to 1, the code will be run sequentially.
Value
A list having two fields:
split_by_resolution: A five-level list. The hierarchy is as follows:
the configuration name: concatenation between the object name provided by the user, the number of neighbors, the graph type and the clustering method
the resolution value \(\gamma\)
the number of clusters k that can be obtained using the specified resolution
a
partitionsfield containing the partitions obtained with resolution \(\gamma\) and have k clusters along with aeccfield that contains the Element-Centric Consistency of the partition listthe structure of a partition, which consists in having a
mbfield with the flat membership vector,freqdenoting its frequency andseed, that is the seed used to obtain this partition in this configuration.
split_by_k: has a similar structure, but the resolution level is removed. The partitions obtained in a configuration with the same number of clusters will be merged into the same list.
Examples
set.seed(2021)
# create an artificial expression matrix
expr_matrix = matrix(runif(500*10), nrow = 500)
# get the PCA embedding of the data
pca_embedding = irlba::irlba(expr_matrix, nv = 2)
pca_embedding = pca_embedding$u %*% diag(pca_embedding$d)
rownames(pca_embedding) = as.character(1:500)
# run the function on the pca embedding
resolution_result = get_resolution_importance(embedding = pca_embedding,
resolution = c(0.8, 1),
n_neigh = c(5, 7),
n_repetitions = 5,
clustering_method = 1,
graph_type = 2,
object_name = "name_example")
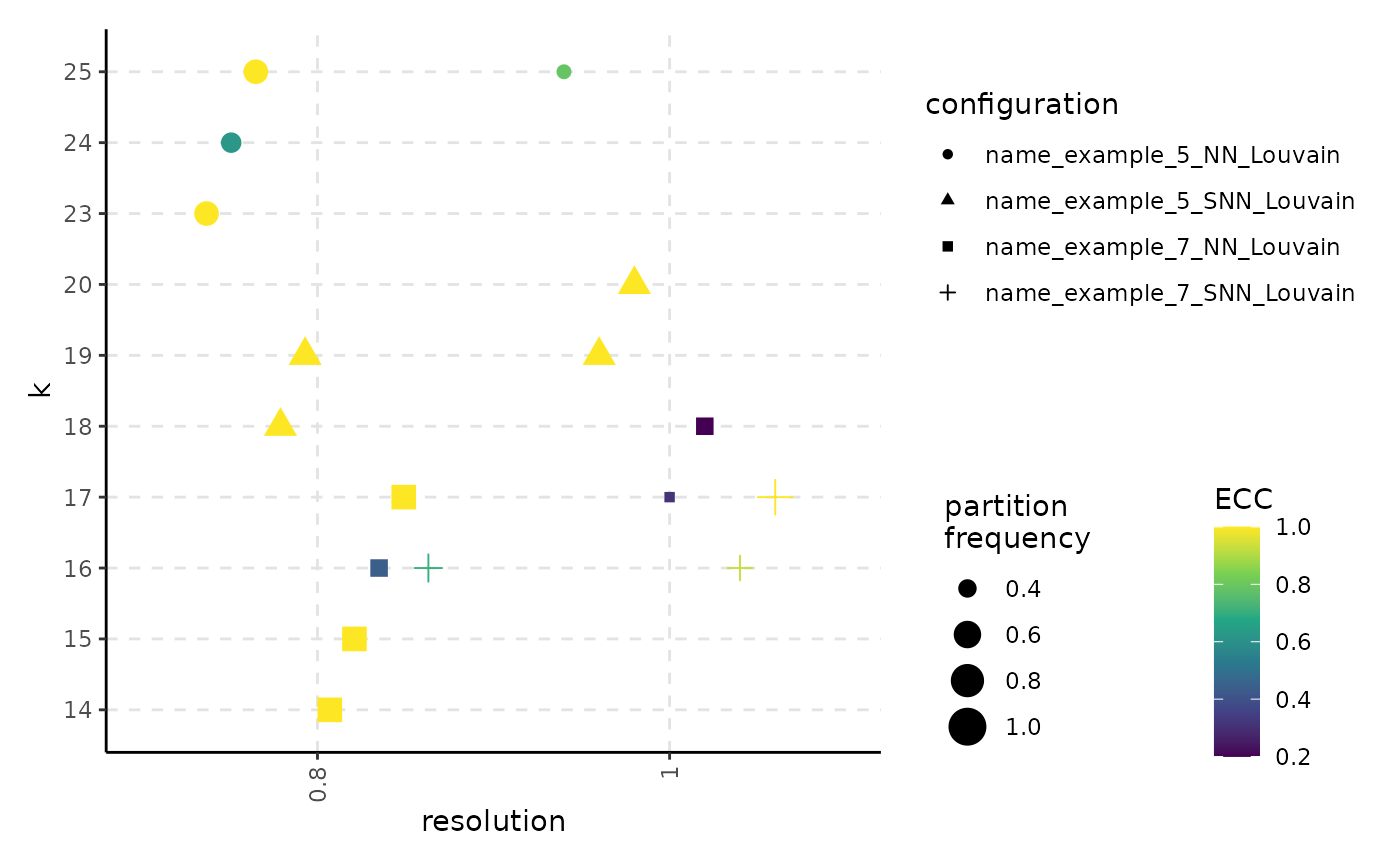
plot_k_resolution_corresp(resolution_result)