Evaluate Feature Set Stability
Source:R/stability-based-parameter-assessment.R
get_feature_stability.RdEvaluate the stability of clusterings obtained based on incremental subsets of a given feature set.
get_feature_stability(
data_matrix,
feature_set,
steps,
feature_type,
n_repetitions = 100,
seed_sequence = NULL,
graph_reduction_type = "PCA",
npcs = 30,
ecs_thresh = 1,
ncores = 1,
algorithm = 4,
...
)Arguments
- data_matrix
A data matrix having the features on the rows and the observations on the columns.
- feature_set
A set of feature names that can be found on the rownames of the data matrix.
- steps
Vector containing the sizes of the subsets; negative values will be interpreted as using all features.
- feature_type
A name associated to the feature_set.
- n_repetitions
The number of repetitions of applying the pipeline with different seeds; ignored if seed_sequence is provided by the user.
- seed_sequence
A custom seed sequence; if the value is NULL, the sequence will be built starting from 1 with a step of 100.
- graph_reduction_type
The graph reduction type, denoting if the graph should be built on either the PCA or the UMAP embedding.
- npcs
The number of principal components.
- ecs_thresh
The ECS threshold used for merging similar clusterings.
- ncores
The number of parallel R instances that will run the code. If the value is set to 1, the code will be run sequentially.
- algorithm
An index indicating which community detection algorithm will be used: Louvain (1), Louvain refined (2), SLM (3) or Leiden (4). More details can be found in the Seurat's
FindClustersfunction.- ...
additional arguments passed to the umap method.
Value
A list having one field associated with a step value. Each step contains a list with three fields:
ecc - the EC-Consistency of the partitions obtained on all repetitions
embedding - one UMAP embedding generated on the feature subset
most_frequent_partition - the most common partition obtained across repetitions
Note
The algorithm assumes that the feature_set is already sorted when performing the subsetting. For example, if the user wants to analyze highly variable feature set, they should provide them sorted by their variability.
Examples
set.seed(2021)
# create an artificial expression matrix
expr_matrix = matrix(c(runif(100*10), runif(100*10, min = 3, max = 4)), nrow = 200, byrow = TRUE)
rownames(expr_matrix) = as.character(1:200)
colnames(expr_matrix) = paste("feature", 1:10)
feature_stability_result = get_feature_stability(data_matrix = t(expr_matrix),
feature_set = colnames(expr_matrix),
feature_type = "feature_name",
steps = 5,
npcs = 2,
n_repetitions = 10,
algorithm = 1,
# the following parameters are used by the umap function and are not mandatory
n_neighbors = 3,
approx_pow = TRUE,
n_epochs = 0,
init = "random",
min_dist = 0.3)
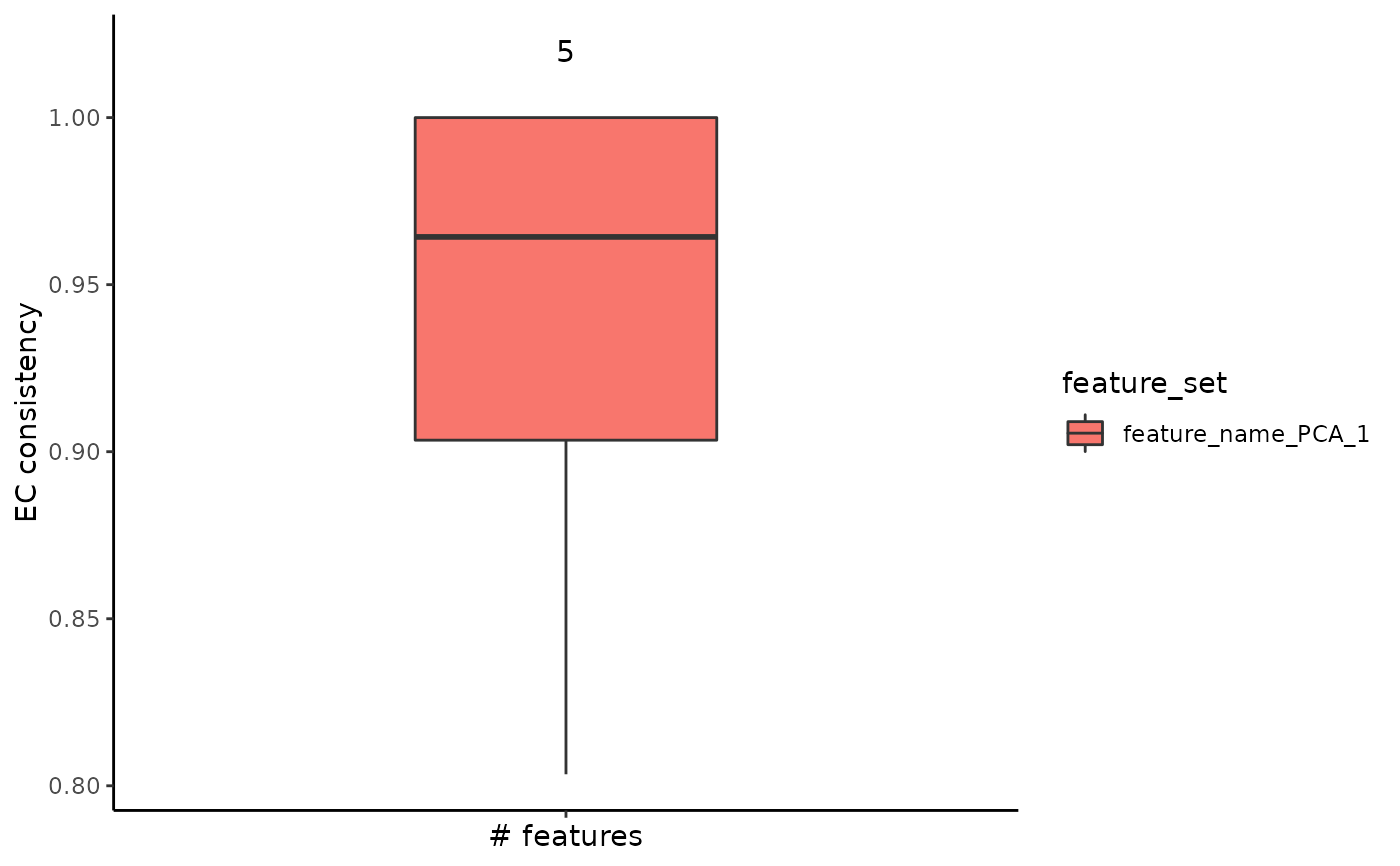
plot_feature_stability_boxplot(feature_stability_result)